Bacterial Genetics Summary By Kazibwe Patrick
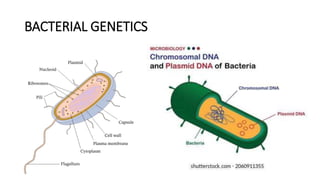
- 2. Bacterial Genetics 1. Microbial Genome(CHROMOSOMAL DNA) - Essential for the organism’s survival and contains genes responsible for various cellular functions. - Replication - Transcription – Translation 2. Plasmid(EXTRACHROMOSOMAL DNA) -Responsible for genetic exchange & variation in bacteria; provide selective advantages when in host, such as antibiotic resistance, toxin production, or metabolic capabilities. - Mutations - Transduction - Conjugation – Transformation - Transposition
- 3. Cell Division Of Bacteria “Binary Transverse Fission” • Cell elongates as growth occurs longitudinal axis • When certain length reached; septum produced in transverse axis midway btn the cell ends NB; DNA replication preceeds septum formation.
- 5. DNA Replication • It is semiconservative type • It begins at the origin of replication oriC; a 245 base pair sequence 1. The oriC opened by DnaA Protein 2. Parent strands unwind by DNA gyrase & @ acts as a template for synthesis of complementary strand. 3. New strands synthesized by DNA polymerase III 4. Process goes bi-directional until replication forks meet; a point at which 2 double stranded daughter DNAs are formed 5. Ends of fully & newly formed strands are joined by DNA ligase to form circular chromosomes
- 6. 1
- 7. Replication fork 1. Single-stranded DNA binding protein - coats single strands preventing them from being denatured 2. DNA gyrase & helicase -Unwind DNA duplex 3. DNA polymerase III - Adds nucleotides to the 3’ end o the leading strand & Okazaki fragments to the RNA primers of the lagging strand 4. DNA polymerase I & ligase - Remove RNA primers replacing them with appropriate DNA segments & join the Okazaki fragments 4. RNA PRIMASE -Adds RNA primers
- 10. Mut SHL Repair System • During replication; base pair mismatch in growing chain are corrected by the 3’ to 5’ exonuclease activity of DNA polymerase III 1. Mut S binds to mismatched base pair 2. Mut H moves along duplex till it finds a methylated base in parent strand 3. Mut L binds to Mut S & Mut H; Mut H nicks the unmethylated strand 4. Helicase & single stranded exonuclease remove the DNA segment 5. Polymerase & ligase add the correct DNA segment in the gap 6. Daughter strand is then methylated
- 13. Transcription • Copy of genes from DNA to RNA (tRNA & rRNA); where they are expressed as proteins needed for sustaining life. • Only short segments of DNA are copied -Assembly of transcription complex - Initiation -Elongation - Termination -Regulation transcription
- 14. 1. Assembly of transcription complex • Only 1 strand (antisense/template/non-coding) of DNA is copied • RNA polymerase activity & sigma factor • RNA polymerase & sigma binds to the promoter site (beginning of a gene) forming a closed promoter complex. • DNA unwinds to form open promoter complex
- 15. 2. Initiation & Elongation • RNA P begins mRNA synthesis by adding a purine nucleotide • After addition of 5 or 6 nucleotides, sigma is released & RNA P continues down the template. "elongation”
- 16. 4.Termination • Rho-independent termination; The G-C rich region with in the transcript forms a hair pin loop Weak pairing of A (DNA template) : U (RNA transcript) Double helix zips up and RNA transcripts dissociates from DNA
- 17. • Rho-dependent; Rho protein(a helicase) binds to C-rich region in transcript & advances in 5’ to 3’ direction till it meets transcription bubble. Bring about unwinding of the transcript & template; and mRNA is released
- 18. Regulation Transcription • Bacteria can adapt to specific environmental conditions by altering levels of mRNA. 1. Negative regulation; The environmental signals interfere with transcription initiation Regulation Of Lac Operon -Genes of lac operon can only be expressed when lactose is the only carbon source; Lac Z, Lac Y, Lac A, (lacZ βgalactosidase, lacY lactose permease, and lacA transacetylase )these are necessary for lactose catabolism - In absence of lactose repressor monomers form tetramer that binds to promoter & blocks transcription - In presence of lactose, it binds & inactivates the repressor
- 20. 2. Positive regulation; environmental signals facilitate transcription Regulation Of Mal Operon - Presence of maltose activates maltose genes - Active Malt gene product binds to promoter of genes involved in the transport & catabolism of maltose
- 21. TRANSLATION • Conversion mRNA to protein • Components; Ribosomes & tRNA • RIBOSOMES; - Consist of both protein and ribosomal RNA (rRNA). - Site where Amino acids are linked together by peptide bonds to form protein -Prokaryotic ribosomes is 70s(svedberg unit) having 30s subunit & 50s subunit
- 23. 1. Initiation 30s subunit binds to Shine-Delgarno Sequence (GGAGGU) on mRNA through its complementary sequence ACCUCC with the help of initiation factors (IF1 & IF3) on the 3’ end of the 16s rRNA GGAGGU is 4-6 nucleotides from the initiation codon (AUG)
- 24. IF2 brings an initiator tRNA charged with initiator amino acid N-formyl- Methionine Pairing occurs between anticodon UAC & complementary codon AUG Larger subunit 50s then binds to the complex & IF1, IF2, IF3 & 16S rRNA are released. A site; entry site for new tRNA charged with amino acrid P site; occupied by tRNA with amino acid of growing polypeptide chain E site; exit for tRNA after delivering amino acid
- 26. 2. Elongation A new tRNA carrying an amino acid enters A site then matching between anticodon of tRNA & codon on mRNA occurs; those with incorrect anticodons are rejected & replaced by new till a right amino-acyl tRNA enters A site A peptide bond made between the 2 adjacent amino acids tRNA in P site releases amino acid/peptide chain to tRNA in A site The ribosome moves to one triplet forward on mRNA; the empty tRNA moves to the E- site A site is now empty & ready to accept new tRNA
- 27. 3. Termination Occurs when a termination codon occupies the A-site and the codon is recognized by a release factor (RF) RF1 recognizes termination codons UAA and UAG, whereas RF2 recognizes UAA or UGA RF3-GTP,( GTPase protein enhances activity of RF1 & RF2) then binds to the ribosome and catalyzes the cleavage of the peptide chain from the last tRNA GTP provides energy for the dissociation of the ribosomes and the release of the RFs
- 28. Basic characteristics of plasmids Extrachromosomal DNA, replication occurs by bacterial cell machinery Circular double stranded DNA Variable size (100 to 1000)bp Copy number: 1-30 copies per cell Some are transferable > conjugation; Only to closely related species except for promiscuous plasmids Plasmid encoded genes not essential to the growth of organism but rather survival.
- 29. • Plasmids encode genes for specialized metabolism Biodegradation of complex organic molecules e.g. some Pseudomonas This allows the bacterium to utilize unusual organic compounds as carbon and energy source
- 30. ORGANISM FACTOR DISEASE Escherichia Coli Enterotoxin Diarrhea Clostridium Tetani Neurotoxin Tetanus Staphylococcus Aureus Coagulase Enterotoxin Food Poisoning/Skin Infections/Boils Streptococcus Mutans Dextransucrase Tooth Decay Agrobacterium Tumefaciens Tumor Crown Gall Staphylococcus Spp Antibiotic Resistance Various Examples Of Virulence Factors Encoded In Plasmids
- 31. Resistance (R) -plasmids • Classified as R-plasmids because of the R-factors that encode antibiotic- resistance determinants • A plasmid can acquire additional R-factors • Implications: R-factors affect therapeutic efficacy Antibiotic sensitive bacterium may become resistant following acquisition of R-plasmids. Neisseria gonorrhoeae has plasmid encoding β-lactamase hence resistance to ampicillin.
- 32. DRUG MECHANISM OF RESISTANCE Beta Lactams Synthesis Of Beta Lactamase Chloramphenicol Synthesis Of Enzyme That Acylates The Drug Aminoglycoside Synthesis Of Enzyme That Inactivate Drug By Acetylation, Phosphorylation, Or Adenylation Tetracycline Synthesis Of Membrane Protein Capable Of Pumping Out Drug Before It Acts On Ribosomes Erythromycin Synthesis Of Enzyme That Methylates 23s Ribosomal RNA TRIMETHOPRIM SYNTHESIS OF MUTANT TRIMETHOPRIM
- 33. GENETIC VARIATION IN BACTERIA • Mechanisms of genetic variation in bacteria: i) Transformation ii) Conjugation iii) Transduction (Generalized & Specialized) iv) Transposition v) Mutation
- 34. DNA Recombination • Natural way by which genetic materials are linked
- 35. Transformation • Bacterium takes in DNA from its environment. This DNA can come from other bacteria that have shed it. • Conversion of avirulent rough (R) type of S. pneumoniae to virulent smooth (S) type by transformation of R type cells with DNA obtained from S type cells
- 36. • Virtually all bacteria have an ability to take up DNA from the environment, provided they are competent • Transformation competence is a state of a bacterial cell during which the usually rigid cell wall can transport a relatively large DNA macromolecule • Some bacteria, such as Hemophilus, Streptococcus, or Neisseria (during a certain stage of cell division) can take up DNA => natural competence. • Treatment of such cells that are usually unable to take up DNA under natural conditions with CaCl2 or RbCl alters their envelope, and they become competent => artificial competence
- 37. Fate of DNA after entering into a competent cell • Transformation of bacteria with a linear DNA fragments is more complex because the new DNA is a natural target for hydrolysis by intracellular enzymes that degrade noncircular DNA. • One mechanism whereby foreign DNA can escape degradation is incorporation into the chromosome of the recipient cell via recombination
- 38. Conjugation • Mechanism of DNA exchange mediated by plasmids • Performed by transmissible plasmids • Donor cells have transmissible plasmids; those cells that receive plasmids recipients • Fertility (F) plasmid of E. coli or the large R plasmids of a variety of other bacteria (self-transmissible plasmids), encode genes responsible for their own replication, maintenance within the bacterial cell and the promotion of DNA transfer via conjugation
- 39. • Transfer functions involve pilus
- 40. Transduction • Is the process of transferring genes between bacteria, which is mediated by bacteriophages (viruses that infect bacteria) • General transduction: random transfer of all bacterial genes at low but identical frequencies • Specialized transduction: certain genes are transferred at very high frequencies, whereas others are transferred at low rates or not at all • Specialized transduction requires incorporation of the viral DNA into the bacterial chromosome, and it is therefore carried out only by lysogenic bacteriophages
- 41. • Specialized transduction results in phage mediated transfer of genes that are near the attachment site of the lysogenic prophage on the chromosome.
- 44. Transposition • An Intracellular gene transfer where a gene moves from one place on the chromosome to another. OR: • A gene moves between a chromosomal site and a plasmid site • The site-specific recombination utilizes DNA sequences that define the site of transposition, and specialized enzymatic machinery that catalyzes the transpositional event • The simplest form of such a mobile gene is an insertion sequence (IS)
- 46. • A common feature of IS: they contain short (16 to 41 bp) inverted repeat sequences at their ends. • The IS also encodes at least one enzyme, called transposase, that specifically mediates the site-specific recombination event during transposition