DNA Traducción
- 3. Fig. 13-4, p. 283 Nontemplate strand Transcription DNA Template strand mRNA (complementary copy of template DNA strand) Codon 1 Codon 2 Codon 3 Codon 4 Codon 5 Codon 6 Polypeptide Met Thr Cys Glu Cys Phe Translation ‘ ‘ ‘ ‘ ‘ ‘
- 6. Fig. 13-20a, p. 299 Normal DNA sequence Normal mRNA sequence Normal protein sequence BASE-SUBSTITUTION MUTATIONS Missense mutation Nonsense mutation (Stop) (Stop) (Stop)
- 7. Fig. 13-20b, p. 299 FRAMESHIFT MUTATIONS Normal DNA sequence Normal mRNA sequence Normal protein sequence (Stop) (Stop)
- 17. Metilación de eF-Tu Fosforilación de elF-2 (eucariotas)
Hinweis der Redaktion
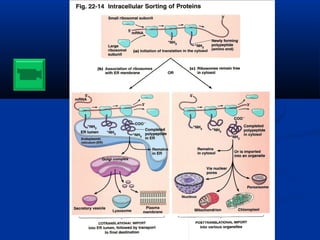
- Figure 22.29 Secretory pathway in eukaryotic cells. Proteins whose synthesis begins in the cytosol are transported into the lumen of the endoplasmic reticulum. After further modification in the Golgi apparatus, the proteins are secreted.
- Figure 13.4: An overview of transcription and translation. In transcription, messenger RNA is synthesized as a complementary copy of one of the DNA strands, the template strand. Messenger RNA carries genetic information in the form of sets of three bases, or codons, each of which specifies one amino acid. Codons are translated consecutively, thus specifying the linear sequence of amino acids in the polypeptide chain. Translation requires tRNA and ribosomes (not shown). The figure depicts transcription and translation in bacteria. In eukaryotes, transcription takes place in the nucleus and translation occurs in the cytosol.
- Figure 22.17 Shine-Dalgarno sequences in E. coli mRNA. (a) Ribosome-binding sites at the 5' end of mRNA for several E. coli proteins. The Shine-Dalgarno sequences (red) occur immediately upstream of initiation codons (blue). (b) Complementary base pairing between the 3' end of 16S rRNA and the region near the 5' end of an mRNA. Binding of the 3' end of the 16S rRNA to the Shine-Dalgarno sequence helps establish the correct reading frame for translation by positioning the initiation codon at the ribosome’s P site.
- Figure 13.20: Mutations. (a) Missense and nonsense mutations are types of base-substitution mutations. A missense mutation results in a polypeptide of normal length, but with an amino acid substitution. A nonsense mutation results in the production of a truncated (shortened) polypeptide, which is usually not functional.
- Figure 13.20: Mutations. (b) A frameshift mutation results from the deletion (shown) or insertion of one or two bases, causing the base sequence following the mutation to shift to a new reading frame. A frame shift may produce a stop codon downstream of the mutation (which would have the same effect as a nonsense mutation caused by base substitution), or it may produce an entirely new amino acid sequence.
- Figure 22.5 Tertiary structure of tRNA. The cloverleaf-shaped molecule shown in Fig. 22.4 actually folds up into this three dimensional shape. The tertiary structure of tRNA results from base pairing between the TC loop and the D loop, and two stacking interactions that (a) align the TC arm with the acceptor arm, and (b) align the D arm with the anticodon arm. For clarity, only the ribose-phosphate backbone is shown here.
- Figure 22.8 Base pairing at the wobble position. The tRNAAla molecule with the anticodon IGC can bind to any one of three codons specifying alanine (GCU, GCC, or GCA) because I can pair with U, C, or A. Note that the RNA strand containing the codon and the strand containing the anticodon are antiparallel.
- Figure 22.15 Sites for tRNA binding in prokaryotic ribosomes. During protein synthesis, the P site is occupied by the tRNA molecule attached to the growing polypeptide chain, and the A site holds an aminoacyl-tRNA. The growing polypeptide chain passes through the tunnel of the large subunit.
- Figure 22.12 Comparison of prokaryotic and eukaryotic ribosomes. Both types of ribosomes consist of two subunits, each of which contains ribosomal RNA and proteins. The large subunit of the prokaryotic ribosome contains two molecules of rRNA: 5S and 23S. The large subunit of almost all eukaryotic ribosomes contains three molecules of rRNA: 5S, 5.8S, and 28S. The sequence of the eukaryotic 5.8S rRNA is similar to the sequence of the 5' end of the prokaryotic 23S rRNA.
- Figure 22.17 Shine-Dalgarno sequences in E. coli mRNA. (a) Ribosome-binding sites at the 5' end of mRNA for several E. coli proteins. The Shine-Dalgarno sequences (red) occur immediately upstream of initiation codons (blue). (b) Complementary base pairing between the 3' end of 16S rRNA and the region near the 5' end of an mRNA. Binding of the 3' end of the 16S rRNA to the Shine-Dalgarno sequence helps establish the correct reading frame for translation by positioning the initiation codon at the ribosome’s P site.
- Figure 22.32 Translocation of eukaryotic proteins into the lumen of the endoplasmic reticulum.